
You never know… a random walk may lead to serendipity.
Today’s xkcd comic is well-timed because just yesterday I sent out an announcement of the availability of my implementation of Cartesian Genetic Programming for ECJ, a Java-based evolutionary computing software framework. Genetic programming (GP) is a problem-solving technique in computer science that is inspired by evolution in biology. You start with a population of randomized computer programs and measure the “fitness” of each program. The fitness is a measurement of how well a program solves a particular problem. You can think of the program itself as the “gene”.
In this simulation of evolution, the best programs in the bunch are selected for “breeding” for the next generations. Breeding is done by exchanging pieces of genetic material – in this case, we exchange pieces of the computer programs themselves. Good programs have different parts that are useful, and these parts are combined or exchanged in new “child” programs that are even better than their parents. Then, every so often, random mutations are introduced into the programs to promote diversity. Good mutations survive to future generations, and bad ones die out.

Parent programs producing offspring by exchanging pieces of themselves. Credit: Michael Adam Lones, Enzyme Genetic Programming
This sounds like a completely random way to solve a problem, but it is surprisingly effective for many kinds of problems, such as learning mathematical expressions that can describe some data set, discovering winning game-playing strategies, making forecasts and predictions from data sets, learning decision trees to classify data, evolving emergent behavior, and optimization of complex systems. Of great interest to me in applying genetic programming is the emergence of unique and fascinating solutions that are not likely to be conceived by human minds.
Cartesian Genetic Programming (CGP), the genetic programming technique I implemented for ECJ, is a variation of genetic programming invented by Julian Miller. CGP uses simple lists of numbers to represent the programs (most GP implementations use some kind of explicit tree representation). It has some interesting benefits over traditional tree-based genetic programs, such as improved search performance, reduced overhead, and less “bloat” in generated programs. My implementation includes some sample problems: regression (fitting a bunch of data to an equation), classification (identifying a species of iris flower based on simple measurements of its parts, or predicting if a breast cancer tumor will be benign or malignant), and parity (counting the number of “on” bits in a binary string).
The iris classification problem is a classic machine learning problem, dating all the way back to 1936. You have a set of measurements taken from various kinds of iris flowers, and your task is to figure out which species it is: iris virginica, iris versicolor, or iris setosa.

From left to right: iris virginica, iris versicolor, and iris setosa.
Starting with randomized programs, one of my CGP tests evolved the following programs (expressions) which correctly classified about 95% of the irises:
virginica = nand (> (- (+ 1.7777395 (/ sepalwidth sepallength)) (if 0.0053305626 petalwidth -0.6896746)) (/ -0.8330147 (neg 1.6308627))) (- (* (+ (- (+ 1.7777395 (/ sepalwidth sepallength)) (- petallength petalwidth)) 1.7777395) (- petallength petalwidth)) petalwidth)
versicolor = * (nand (> (- (+ 1.7777395 (/ sepalwidth sepallength)) (if 0.0053305626 petalwidth -0.6896746)) (/ -0.8330147 (neg 1.6308627))) (- (* (+ (- (+ 1.7777395 (/ sepalwidth sepallength)) (- petallength petalwidth)) 1.7777395) (- petallength petalwidth)) petalwidth)) (- (+ 1.7777395 (/ sepalwidth sepallength)) (- petallength petalwidth))
setosa = - (+ 1.7777395 (/ sepalwidth sepallength)) (- petallength petalwidth)
I don’t expect you to be able to read and understand the expressions – they are in a format that isn’t easy to read! What’s more important is that I didn’t have to create them myself – the CGP algorithm discovered them for me using only the input data (iris measurements) and the fitness function (a measurement of how many irises were correctly identified).
CGP also had good results when I tested the Wisconsin breast cancer data set. This data set contains measurements taken from fine needle biopsies of suspicious breast lumps. Our task is to predict whether the lumps are benign or malignant using only the measurements.
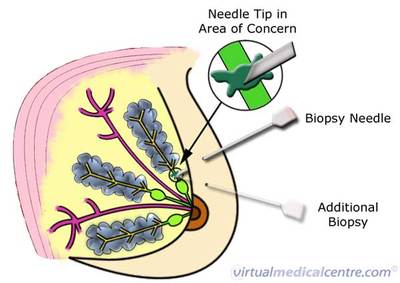
Fine needle aspiration. A thin needle is used to sample material from a suspicious lump in a breast. The data set contains the microscopic measurements taken from the sampled material.
CGP evolved the following program that correctly diagnoses the tumors 95% of the time:
malignant = not (nor (+ (+ (* cellShapeUniformity 1.5756812) bareNuclei) (<= (> (= cellSizeUniformity 0.08695793) -1.9803793) normalNucleoli)) (nor (> (- (iflez (if blandChromatin mitoses 0.75769496) bareNuclei normalNucleoli) (* 1.97491 (+ cellSizeUniformity clumpThickness))) (> 0.08695793 singleEpiCellSize)) (nor (>= marginalAdhesion (not (- (= (> (or clumpThickness 1.5756812) (= cellSizeUniformity 0.08695793)) (+ cellSizeUniformity clumpThickness)) (* (* 1.97491 (+ cellSizeUniformity clumpThickness)) mitoses)))) (iflez (if blandChromatin mitoses 0.75769496) bareNuclei normalNucleoli))))
Again, using only a fitness function and some test data, we are able to evolve a highly accurate cancer diagnosis tool.
Seems too good to be true? Well, it can be. Overfitting is a big problem when evolving these kinds of classifiers. For example, if each of the malignant tumor patients happened to be wearing red shoes during the biopsy (and the shoe type was included in the data set), a machine might be inclined to think that wearing red shoes was what determined the diagnosis. So, the evolved classifier is going to be very sensitive to whatever data set you unleash it upon.
Oh and then there’s the whole “no free lunch” thing. But that’s a depressing topic for another time.
If you want to find out more about my CGP implementation, check out the documentation. If you want to give it a whirl yourself, grab the distribution (you need at least Java 1.5). Check out the ECJ project page for more info about the evolutionary software framework my CGP implementation uses.

Pingback: ideonexus.com » Blog Archive » Patterns in the PHP Random Function